MLZ is a cooperation between:
 > Technische Universität München
> Technische Universität München > Helmholtz-Zentrum Hereon
> Helmholtz-Zentrum Hereon
 > Forschungszentrum Jülich
> Forschungszentrum Jülich
MLZ is a member of:
 > LENS
> LENS > ERF-AISBL
> ERF-AISBL
MLZ on social media:

MLZ (eng)
Lichtenbergstr.1
85748 Garching
14.11.2016
Improved Insight into the Structure of Lipid Membranes: Combining the Strengths of Neutron Reflectivity and Molecular Dynamics
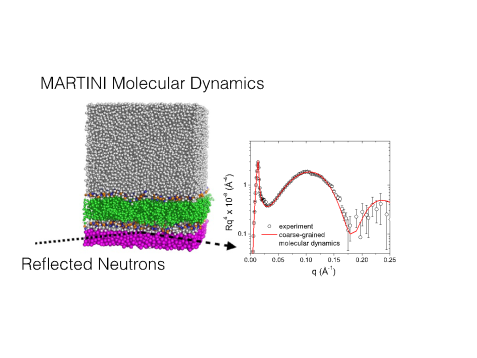
A typical molecular dynamics snapshop and comparison with reflectivity experiments. © Forschungszentrum Jülich
Neutron reflectometry is one of the most suitable methods for investigating the structure of lipid membranes in model systems. These model systems are more easily prepared and experimentally controlled than natural lipid membranes, which are an important integral part of animal and plant cells. A scientist from the Jülich Centre for Neutron Science has now presented a method which delivers more precise results and should pave the way for studying membranes of a more complex nature. The method combines the advantages of neutron reflectometry and molecular dynamic simulations.
Biological lipid membranes represent an important self-assembly system in living organisms. In their most abundant form as the primary constituent of the cell membrane, they provide a bilayer fluid matrix for embedded membrane proteins that in turn regulate the cell’s communication with the surrounding environment and its transport of chemical species. Biomembranes are immensely complex containing mixtures of many different types of lipids, cholesterol and proteins. Usually in biophysical studies, simpler counterparts are used, consisting of a single or a few different lipid types.
The existence of methods that permit the reconstitution of model biomembranes on flat solid/liquid interfaces, facilitates the use of a multitude of surface analytical experimental techniques for the study of their structure and dynamics. Among these techniques, neutron reflectometry is probably the most powerful probe of their structure. The special nature of neutrons enables them to penetrate deeply into many materials, making them ideal probes for buried interfaces, as membranes at the solid/liquid interface. Furthermore, the ability to manipulate contrast by the use of deuterated solvent or lipids offers many possibilities for the extraction of structural information at the sub-nanometer level.
Traditionally, reflectivity data are compared to theoretical curves of simplified models consisting of stratified layers representing the hydrophilic parts, that is the lipid heads, and the hydrophobic parts – the lipid tails – of the membrane system. The end result is a lateral averaged scattering length density profile that gives hints about the molecular distribution across the membrane. In the present work, we attempt to improve the interpretation of reflectivity data by the combined use of molecular simulations and neutron reflectivity.
A set of neutron reflectivity measurements was acquired at the MARIA reflectometer, using 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC)-supported membranes on silicon substrates. This exact system was simulated using a coarse-grained molecular dynamics approach based on the MARTINI force field. Then the system parameters leading to an agreement between simulation and experiment were identified. Results indicate that by carefully tuning coarse-grained simulations, we are able to reproduce the experimental behavior of the system both in the fluid and gel lipid phase. An analysis of the molecular dynamics trajectories offers hints on the structural and dynamical details of the lipid systems. Finally the successful application of this method paves the way for the investigation of more elaborate systems.
A large part of the “Soft Matter/Biology” experiments that take place on the MARIA instrument at the Heinz Maier-Leibnitz Zentrum (MLZ) in Garching attempt to exploit the resolving power of neutron reflectivity for the study of model biological membranes. In the near future, we envision the use of the approach presented here for the preparation of reflectivity experiments and also for the elucidation of subtle structural features in experiments where membranes interact with bio-molecules, e.g. proteins, small peptides, and nanoparticles that are candidates for drug delivery applications.
Originalpublikation:
Alexandros Koutsioumpas; Combined Coarse-Grained Molecular Dynamics and Neutron Reflectivity Characterization of Supported Lipid Membranes; J. Phys. Chem. B 120 (44), 11474 (2016)
MLZ is a cooperation between:
 > Technische Universität München
> Technische Universität München > Helmholtz-Zentrum Hereon
> Helmholtz-Zentrum Hereon
 > Forschungszentrum Jülich
> Forschungszentrum Jülich
MLZ is a member of:
 > LENS
> LENS > ERF-AISBL
> ERF-AISBL
MLZ on social media:



